14. Metropolis coupled Markov chain Monte Carlo [(MC)3]¶
The Markov chain Monte Carlo (MCMC) method implemented in BAMM estimates the posterior probability distribution of diversification models that best describe a particular phylogenetic dataset. Before version 2.0.0, BAMM used a single Markov chain to explore the landscape of models and their parameter values. A single chain, however, may get stuck in local optima, which results in less mixing and more time needed for convergence.
Metropolis coupled Markov chain Monte Carlo [(MC)3], introduced in BAMM 2.0.0, increases the number of Markov chains that explore the landscape of models and their parameters. Not only are there more chains, but each one has a different temperature. The cold chain is the main chain and behaves as before. The heated chains explore a landscape that is flatter than the landscape explored by the cold chain. Therefore, it is easier for a heated chain to cross deep valleys in the landscape and not get stuck in local optima.
After the chains independently explore the landscape for a certain number of generations, a proposal is made to swap the temperatures of two random chains. If a swap involving the cold chain is successful, it will cause a previously heated chain to become the main chain. As a result, a chain that is stuck in a local optimum may immediately jump to another area of the landscape.
The implementation of (MC)3 in BAMM is described in
Shi & Rabosky 2015.
For each chain 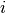 , its temperature
, its temperature 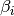 is set to
is set to
![\beta_i = [1 + \Delta T \times (i - 1)]^{-1}](_images/math/6a26ca0b490f6c5714248995e167f3d9704873a5.png) ,
where
,
where 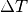 is the temperature increment parameter.
For example, if there are 4 chains and
is the temperature increment parameter.
For example, if there are 4 chains and 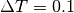 ,
the temperatures of each chain are 1, 0.9091, 0.8333, and 0.7692.
The temperature of the cold chain is always 1.
Note that the value of
,
the temperatures of each chain are 1, 0.9091, 0.8333, and 0.7692.
The temperature of the cold chain is always 1.
Note that the value of 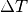 should be greater than 0
and chosen such that the probability of accepting a swap
is between 20% and 60% (Altekar et al. 2004).
should be greater than 0
and chosen such that the probability of accepting a swap
is between 20% and 60% (Altekar et al. 2004).
The temperature of each chain 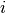 goes into the calculation
of the acceptance probability for a within-model proposal
(i.e., not involving changes in the dimensionality of the model):
goes into the calculation
of the acceptance probability for a within-model proposal
(i.e., not involving changes in the dimensionality of the model):
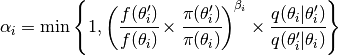
where 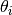 and
and 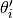 are parameter vectors
corresponding to the current and proposed states for chain
are parameter vectors
corresponding to the current and proposed states for chain 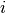 ,
,
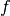 and
and 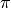 are the corresponding likelihood
and prior density functions,
and
are the corresponding likelihood
and prior density functions,
and 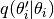 is the relative probability
of proposing a move to parameter vector
is the relative probability
of proposing a move to parameter vector 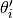 given that the current state is
given that the current state is 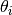 .
A similar calculation is done for the acceptance probability for proposals
that change the dimensionality of the model.
.
A similar calculation is done for the acceptance probability for proposals
that change the dimensionality of the model.
After a certain number of generations, two randomly chosen chains
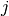 and
and 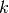 are swapped with acceptance probability
are swapped with acceptance probability
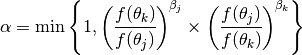
14.1. Multi-core Metropolis coupled MCMC [(MC)4]¶
Using a single CPU, the amount of time a run takes to finish scales linearly with the number of chains. Because chains are mostly independent from each other, except when two chains are chosen to swap states, they may be set up to run on different CPUs in parallel. BAMM implements this parallelization using threads in C++11.
14.2. (MC)3 settings in BAMM¶
14.2.1. Control file settings¶
There are four settings in BAMM that control the behavior of (MC)3.
They are in the template and example control files
under the heading METROPOLIS COUPLED MCMC.
The number of Markov chains to use is specified with numberOfChains.
The 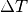 value is specified with
value is specified with deltaT.
The number of generations between chain swap proposals
is specified with swapPeriod.
BAMM will output to a file (chain_swap.txt by default)
the generation in which a swap proposal occurred,
the ranking of the two chains chosen
(1 is the cold chain, 2 is the first heated chain, etc.),
and whether the swap was accepted.
The file to which to output these data is specified with chainSwapFileName.
Our preliminary tests show that four chains produces good MCMC mixing.
The 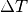 value should be set such that the probability
of accepting a chain swap proposal is between 20% and 60%.
For small to medium sized trees (< 1,000 taxa),
we have found that
value should be set such that the probability
of accepting a chain swap proposal is between 20% and 60%.
For small to medium sized trees (< 1,000 taxa),
we have found that 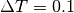 works well.
For large trees,
works well.
For large trees, 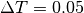 or
or 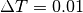 works better.
We have not examined the effects of the swap period in detail,
but in terms of run time,
the smaller the swap period, the longer the run takes to complete.
A swap period of 1000 has worked for us.
works better.
We have not examined the effects of the swap period in detail,
but in terms of run time,
the smaller the swap period, the longer the run takes to complete.
A swap period of 1000 has worked for us.
14.2.2. Determine the best settings¶
In the tools directory of the BAMM GitHub repository,
we have provided an R script, chainSwapPercent.R,
and a bash script (OS X and Linux only), chain-swap-percent.sh,
to help determine the optimal 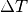 value for a specific data set.
These scripts print out the percent acceptance of the chain swap proposals
by testing all combinations of the given values of
swap period,
value for a specific data set.
These scripts print out the percent acceptance of the chain swap proposals
by testing all combinations of the given values of
swap period, 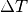 , and number of chains.
Both scripts work similarly,
so choose one or the other depending on convenience.
, and number of chains.
Both scripts work similarly,
so choose one or the other depending on convenience.
To use either of the scripts,
first make sure you are in the directory containing your data files.
We assume that the scripts are located in ~/bamm/tools
and that the BAMM executable can be run as bamm.
The following examples assume that you would like to test
all combinations for the number of chains of 2, 4, and 6,
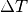 of 0.01 and 0.05, and swap period of 100 and 1000.
To use the R script, run in R:
of 0.01 and 0.05, and swap period of 100 and 1000.
To use the R script, run in R:
source("~/bamm/tools/chainSwapPercent.R")
chainSwapPercent(bammPath = 'bamm', controlfile = 'divcontrol.txt',
nChains = c(2, 4, 6), deltaT = c(0.01, 0.05),
swapPeriod = c(100, 1000), nGenerations = 20000,
burnin = 0.2, deleteTempFiles = TRUE)
This will produce the following output:
|==================================================================| 100%
nChains deltaT swapP percent accepted proposed
1 2 0.01 100 0.98125 157 160
2 2 0.01 1000 1.00000 16 16
3 2 0.05 100 0.85000 136 160
4 2 0.05 1000 0.87500 14 16
5 4 0.01 100 0.98750 158 160
6 4 0.01 1000 1.00000 16 16
7 4 0.05 100 0.88750 142 160
8 4 0.05 1000 0.93750 15 16
9 6 0.01 100 0.98125 157 160
10 6 0.01 1000 1.00000 16 16
11 6 0.05 100 0.75625 121 160
12 6 0.05 1000 0.93750 15 16
These results may be saved into a variable by assigning the variable the result of the chainSwapPercent function.
To use the bash script, run in a terminal:
~/bamm/tools/chain-swap-percent.sh --numberOfChains 2 4 6
--deltaT 0.01 0.05 --swapPeriod 1000 --run bamm -c divcontrol.txt
The output will be similar to that produced by the R script.
